DNA to Protein Converter using Python 3 and Django 2.1
DNA is present in every cell of human being. Informations gathered from DNA can be very usefule in crime investigations, health care and medicine industries.
Here, i have explained how can we produce protin structure from any given DNA sample. Let me give you a quick information about how does DNA sample will look like which we are going to use here and what information from biotech will we be using to produce protin structure.
DNA:
DNA sample will be one dimensional string of characters with four characters to choose from. These characters are A, C, G, and T. They stand for the first letters with the 4 nucleotides used to construct DNA. The full names of these nucleotides are Adenine, Cytosine, Guanine, and Thymine. Each unique three character sequence of nucleotides, sometimes called a nucleotide triplet, corresponds to one amino acid. The sequence of amino acids is unique for each type of protein and all proteins are built from the same set of just 20 amino acids for all living things.
Protin structures:
Protin sturctures that we want to produce will also be of a 1 dimensional string and it will consist values from biotech.
Knowledge from biotech:
To produce, Protin structures from DNA sample we'll use a table. This table provides information about what should be the protein structure if DNA strcuture is this.
So the first step is to read instructions in the DNA and first transcribed them into RNA and the RNA is then translated into proteins.
Here is how our table will look like:
table = {
'ATA':'I', 'ATC':'I', 'ATT':'I', 'ATG':'M',
'ACA':'T', 'ACC':'T', 'ACG':'T', 'ACT':'T',
'AAC':'N', 'AAT':'N', 'AAA':'K', 'AAG':'K',
'AGC':'S', 'AGT':'S', 'AGA':'R', 'AGG':'R',
'CTA':'L', 'CTC':'L', 'CTG':'L', 'CTT':'L',
'CCA':'P', 'CCC':'P', 'CCG':'P', 'CCT':'P',
'CAC':'H', 'CAT':'H', 'CAA':'Q', 'CAG':'Q',
'CGA':'R', 'CGC':'R', 'CGG':'R', 'CGT':'R',
'GTA':'V', 'GTC':'V', 'GTG':'V', 'GTT':'V',
'GCA':'A', 'GCC':'A', 'GCG':'A', 'GCT':'A',
'GAC':'D', 'GAT':'D', 'GAA':'E', 'GAG':'E',
'GGA':'G', 'GGC':'G', 'GGG':'G', 'GGT':'G',
'TCA':'S', 'TCC':'S', 'TCG':'S', 'TCT':'S',
'TTC':'F', 'TTT':'F', 'TTA':'L', 'TTG':'L',
'TAC':'Y', 'TAT':'Y', 'TAA':'_', 'TAG':'_',
'TGC':'C', 'TGT':'C', 'TGA':'_', 'TGG':'W',
}
Conversion of DNA to protin is done on the basis on values corresponding to table. We'll break down DNA sample in pieces of 3 Characters and look for corrsponding value into table. Here is how the function will look like.
def generate_protin(seq):
table = {
'ATA':'I', 'ATC':'I', 'ATT':'I', 'ATG':'M',
'ACA':'T', 'ACC':'T', 'ACG':'T', 'ACT':'T',
'AAC':'N', 'AAT':'N', 'AAA':'K', 'AAG':'K',
'AGC':'S', 'AGT':'S', 'AGA':'R', 'AGG':'R',
'CTA':'L', 'CTC':'L', 'CTG':'L', 'CTT':'L',
'CCA':'P', 'CCC':'P', 'CCG':'P', 'CCT':'P',
'CAC':'H', 'CAT':'H', 'CAA':'Q', 'CAG':'Q',
'CGA':'R', 'CGC':'R', 'CGG':'R', 'CGT':'R',
'GTA':'V', 'GTC':'V', 'GTG':'V', 'GTT':'V',
'GCA':'A', 'GCC':'A', 'GCG':'A', 'GCT':'A',
'GAC':'D', 'GAT':'D', 'GAA':'E', 'GAG':'E',
'GGA':'G', 'GGC':'G', 'GGG':'G', 'GGT':'G',
'TCA':'S', 'TCC':'S', 'TCG':'S', 'TCT':'S',
'TTC':'F', 'TTT':'F', 'TTA':'L', 'TTG':'L',
'TAC':'Y', 'TAT':'Y', 'TAA':'_', 'TAG':'_',
'TGC':'C', 'TGT':'C', 'TGA':'_', 'TGG':'W',
}
protein =""
for i in range(0, len(seq), 3):
try:
codon = seq[i:i + 3]
protein+= table[codon]
except KeyError:
return protein
return protein
We'll put this function into a file.
Using Django to provide web interface
We'll create a Django project
django-admin startproject dna25
then we'll create an app inside the project called dmain
python manage.py startapp dmain
inside this app we'll put the dna_analyzer file and create url structure and view for the home page.
on Homepage we'll create a form and we'll handle the POST request of form in view.
On post method, we'll get the input and call protein_generator function from dna_analyzer file and show the output.
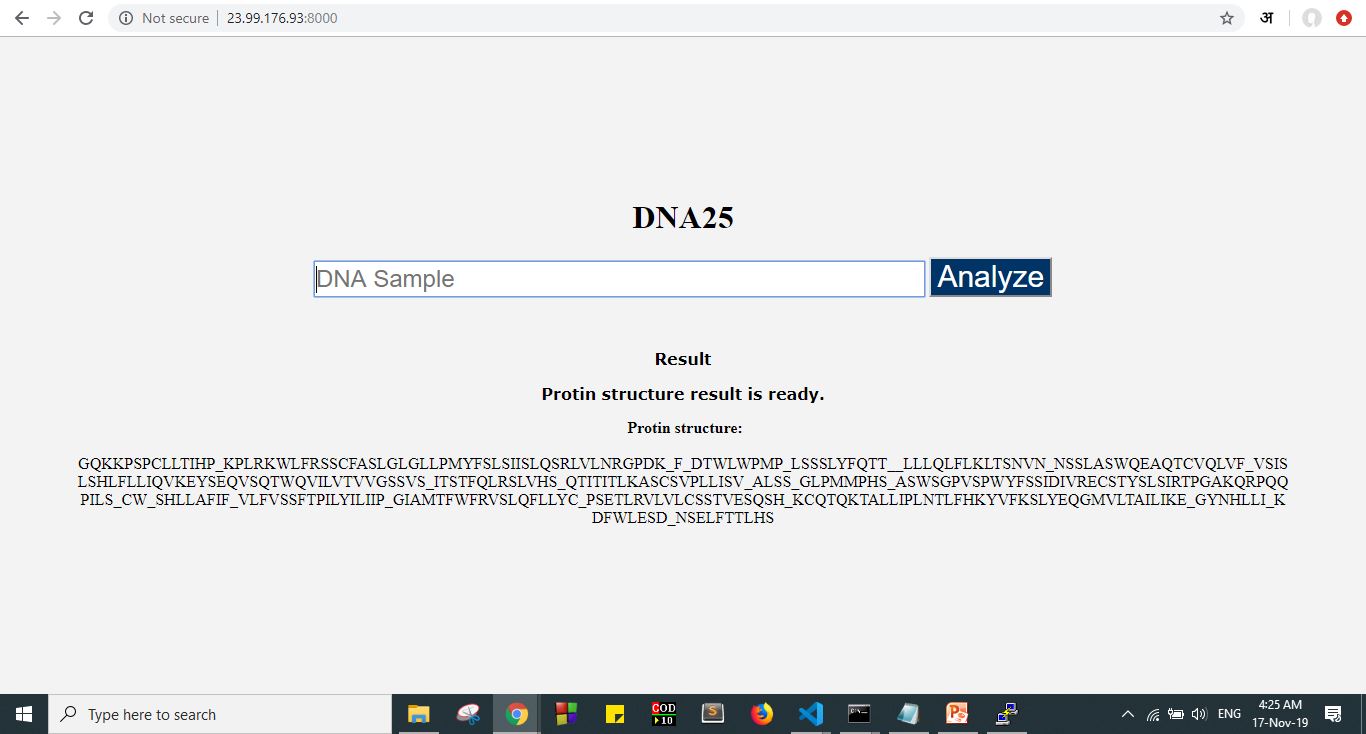
here is how the UI will look like:
A live demo can be seen from here : https://dna25.azurewebsites.net/
I created it during a hackahton at Microsoft.